by Yen Bui
supervisor: Dr. Jeremy Schmit
Kansas State University Physics Department REU Program
This program is funded by the National Science Foundation through grant number PHY-1157044. Any opinions, findings, and conclusions or recommendations expressed in this material are those of the author(s) and do not necessarily reflect the views of the National Science Foundation.
Welcome to my webpage. This page summarizes my experience doing research for the Summer 2012 in the lab of Dr. Jeremy Schmit. I am trying study the general interactions between proteins by studying them in their crystal form in order to understand their patterns of aggregation and solubility.
Below, I describe the Project Overview and my Research Strategy. I have also posted my Final Presentation. Scroll all the way down to learn more About Me.
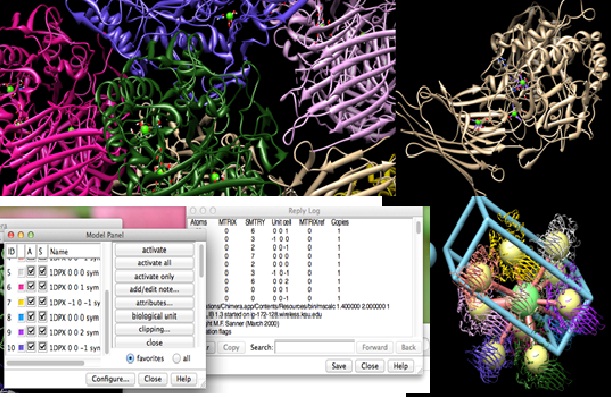
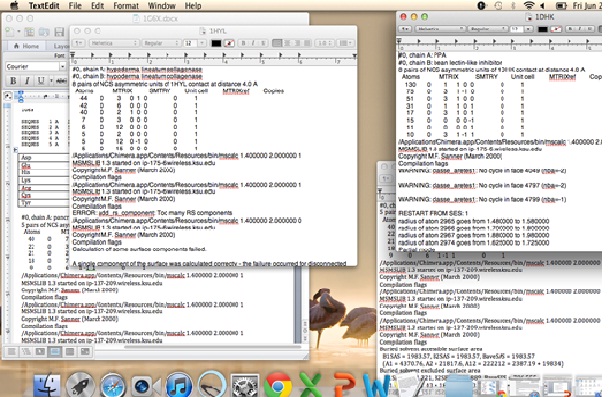
Here is screen shot of a typical screenshot I’ll view.
As reseachers further investigate into the field of pharmacology and biochemistry, protein-protein interactions become of great importance in terms of understanding the various ways in which drug synthesis and delivery will affect the human physiology and enzymatic function. However, as drugs and therapies are continually researched and developed, it becomes very obvious that the basic interactions between proteins are not well understood – especially the ways in which they crystallize. However, it is known that the interactions depend on the fundamental chemistry between residues and the various structures of the protein and its aqueous environment (such as temperature, pH, etc.). Going off the basis of previous research, I worked with Dr. Schmit to further develop his theoretical model describing these protein-protein interactions (dependent upon pH, salt concentration, and temperature) in order to explain how the various bond strengths form within proteins. I did this by using a variety of computational functions to analyze the intermolecular contact points within the protein and explain their tendency to crystallize. This information will give us further insight on why proteins interact the way they do, which will further add to the general knowledge of protein studies – specifically the roles of short-range hydrophobic interactions and long range electrostatic forces between proteins.
Using experimental data we attempted to fit the calculated Enthalpy and Entropy values of various proteins to Dr. Schmit’s theoretical model describing protein interactions.
According to basic protein fundamentals, there are pieces of electrostatics and non-electrostatics that account for the tendency of a protein to aggregate or be repulsed by another. In this project we attempt to account for all those factors using the program:

We used this program to model our proteins in order to calculate the Buried Surface Area (to calculate the Hydrophobic Effect) and Hydrogen-Bonding which would determine the extent of their contribution to the stability of the protein crystals.
However, in order to do this, we had to calculate the electrostatics of the proteins using the information obtained from Chimera and a number of previously published research on our proteins using:
Once the pieces were calculated they were compiled in Microsoft Excel for each protein and analyzed.
Research Progress: I have documented my progress through the various powerpoints presentation I gave:
Final
Presentation:
Click here to download my presentation in powerpoint and pdf formats:
My name is Yen Bui and I am an undergraduate student at Rockhurst University (a small liberal arts school) pursuing a bachelor of science in Chemistry with a minor in biology. As student in general, I am well-rounded in all areas of the physical sciences and have learned to incorporate each subject as a component of grander whole. My approach to acquiring my BS is to gain firm grounding in all of my basic classes and then attempt to apply them in my undergraduate research and eventually work on getting myself a PhD.
My own personal interests are in biomaterials and novel compounds. Although I am a student at Rockhurst University, I sought out and found for myself a professor to work with at a neighboring institute when I realized none of the faculty at Rockhurst were conducting research whose projects were in line with my own. I am currently working in a laboratory at the University of Kansas City, Missouri as a volunteer lab assistant to Dr. Ekaterina Kadnikova. Dr. Kadnikova’s focus is on bioorganic and materials chemistry with an emphasis on enantioselective catalysis, and design of functional polymers and hybrid organic-inorganic materials for biomedical applications. In her lab, I help synthesize nanoparticles with core-shell morpology in which the polymer shell is grown from the inorganic core using enzymatic polymerization.
I saw this REU program at KSTATE as a way to gain invaluable knowledge and experience for all of my future research projects. I had a strong background in chemistry and biology and wanted to become more inter-disciplinary and engage myself in a field more focused on physics. I was able to do that with Dr. Schmit (a theoretical biophysicist) and was also able to gain an opportunity to introduce myself to faculty investigators while visiting and learning from other research groups studying the field of soft matter.
My Research group's home page: Dr. Schmit's Research Group