Research Projects
Large systems often show phenomena that are difficult to anticipate based on the properties of the individual components. Examples of such "emergent properties" include Cooper Pairs emerging from phonons and electron bands in superconductors, the hydrophobic effect emerging from large numbers of water molecules, and political parties emerging from large groups of people. Identifying emergent properties allows us to understand complicated systems by teaching us which details at small scales are important for how the collective system behaves. For example, the emergence of valence electrons from a collection of atomic orbitals allows us to understand molecular structures without solving the complicated quantum problem of interacting electrons. Research in Schmit group focuses on identifying emergent properties and simplifying concepts in large systems of biomolecules. This theme encompasses projects on membraneless organelles, aggregation diseases, self-assembly, drug formulations, and regulatory networks in cells. We primarily used analytic theory (i.e., pen-and-paper theory) and work closely with many experimental groups.
Biological phase transitions
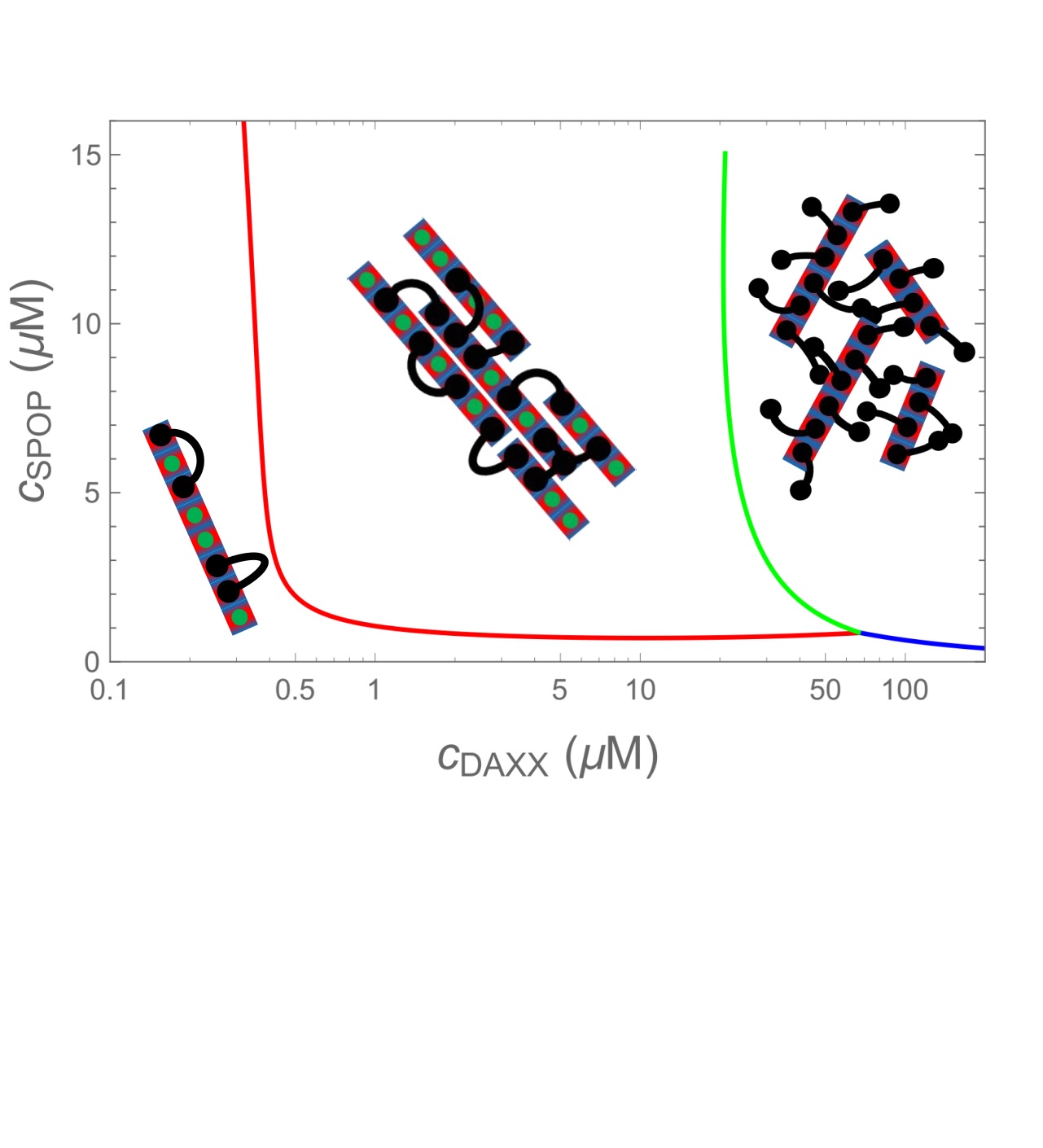 There is a growing list of cellular structures are formed by the spontaneous phase separation of biomolecules. These structures have many common features, such as the presence of multivalent scaffold molecules. However, they also perform many different functions within the cell and have evolved to optimize these specific functions. We are studying how structure/function properties propagate from the nanometer-scale characteristics of the individual molecules to the characteristics of a macroscopic phase.
There is a growing list of cellular structures are formed by the spontaneous phase separation of biomolecules. These structures have many common features, such as the presence of multivalent scaffold molecules. However, they also perform many different functions within the cell and have evolved to optimize these specific functions. We are studying how structure/function properties propagate from the nanometer-scale characteristics of the individual molecules to the characteristics of a macroscopic phase. Protein crystallization
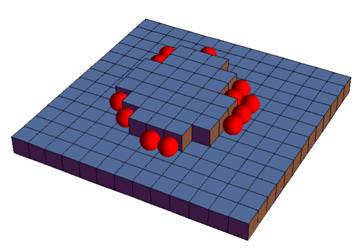 X-ray crystallography is the workhorse technique for determining the three-dimensional structures of folded proteins. Typically the slowest step in this process is the growing of the crystals, which is currently accomplished by high throughput screening (i.e. trial-and-error). Crystallization is a "Goldilocks" problem: the proteins must stick together strongly enough to stabilize the crystal, but not so strongly that the aggregation is uncontrolled. To understand this we need two things. First, we need to understand the forces that make proteins stick together, and how these forces are affected by experimental variables like temperature, pH, and salts. Secondly, we need to understand the kinetic pathways that lead to crystallization and the pathways that lead to non-crystalline aggregates and gelation.
X-ray crystallography is the workhorse technique for determining the three-dimensional structures of folded proteins. Typically the slowest step in this process is the growing of the crystals, which is currently accomplished by high throughput screening (i.e. trial-and-error). Crystallization is a "Goldilocks" problem: the proteins must stick together strongly enough to stabilize the crystal, but not so strongly that the aggregation is uncontrolled. To understand this we need two things. First, we need to understand the forces that make proteins stick together, and how these forces are affected by experimental variables like temperature, pH, and salts. Secondly, we need to understand the kinetic pathways that lead to crystallization and the pathways that lead to non-crystalline aggregates and gelation. Self-assembly
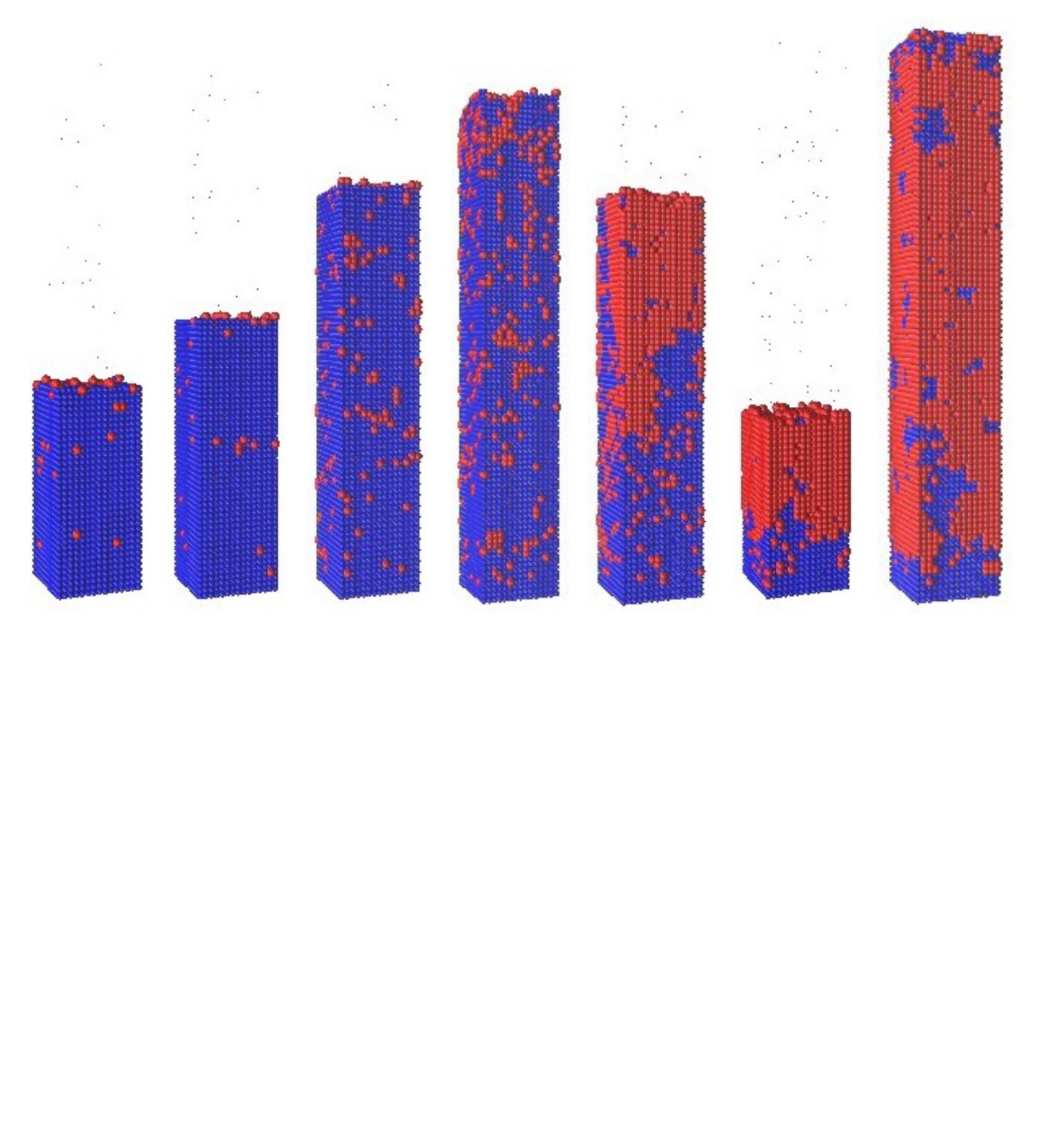 Most biological machines are built by "bottom-up" processes, where building blocks spontaneously assemble into a final product. The complexity of machines observed in nature speaks volumes of the potential of bottom-up assembly in nano-manufacturing and medicine. Experimental groups have made tremendous progress recently in designing nanoscale building blocks with novel shapes and binding affinities. However, in order to mimic the complexity of assembled objects found in nature, we need a better understanding of the kinetic factors that lead to either successful assembly or undesirable aggregates.
Most biological machines are built by "bottom-up" processes, where building blocks spontaneously assemble into a final product. The complexity of machines observed in nature speaks volumes of the potential of bottom-up assembly in nano-manufacturing and medicine. Experimental groups have made tremendous progress recently in designing nanoscale building blocks with novel shapes and binding affinities. However, in order to mimic the complexity of assembled objects found in nature, we need a better understanding of the kinetic factors that lead to either successful assembly or undesirable aggregates. Amyloid fibrils
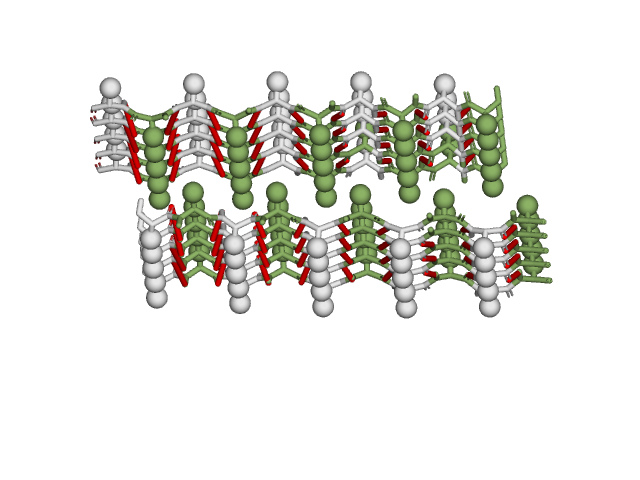 Many diseases, such as Alzheimer's, Type II diabetes, and Mad Cow, are associated with the deposition of fibrillar protein aggregates called "amyloids". These aggregates have a striking degree of morphological similarity despite little sequence homology in the precursor proteins. While a few cases of these diseases are genetic, the large majority are sporadic. What is is that determines whether aggregation begins in middle age, late in life, or not at all? Another complication is the growing evidence that disease progression is driven, not by the fibrils, but by soluble oligomer states. Are these oligomers precursors to fibril formation, or off-pathway aggregates? The goal of this project is to understand the physics of amyloid aggregation in order to develop strategies to remove protein from the most toxic states.
Many diseases, such as Alzheimer's, Type II diabetes, and Mad Cow, are associated with the deposition of fibrillar protein aggregates called "amyloids". These aggregates have a striking degree of morphological similarity despite little sequence homology in the precursor proteins. While a few cases of these diseases are genetic, the large majority are sporadic. What is is that determines whether aggregation begins in middle age, late in life, or not at all? Another complication is the growing evidence that disease progression is driven, not by the fibrils, but by soluble oligomer states. Are these oligomers precursors to fibril formation, or off-pathway aggregates? The goal of this project is to understand the physics of amyloid aggregation in order to develop strategies to remove protein from the most toxic states.Drug formulations
 Protein-based pharmaceuticals are the fastest growing segment of the drug discovery industry. These products are very well tolerated and can offer remarkable specificity. However, proteins are fragile molecules and there are numerous stresses involved with manufacturing, transportation, storage, and delivery. The final formulation must be stable against denaturation and/or aggregation. At the same time the formulation must be compatible with the delivery mechanism. Concentrated solutions are cheaper and easier to administer, but at high concentrations proteins are extremely viscous. This project is a collaboration with Amgen to understand the aggregation and viscosity of protein solutions in order to develop formulations that are both stable and easily administered.
Protein-based pharmaceuticals are the fastest growing segment of the drug discovery industry. These products are very well tolerated and can offer remarkable specificity. However, proteins are fragile molecules and there are numerous stresses involved with manufacturing, transportation, storage, and delivery. The final formulation must be stable against denaturation and/or aggregation. At the same time the formulation must be compatible with the delivery mechanism. Concentrated solutions are cheaper and easier to administer, but at high concentrations proteins are extremely viscous. This project is a collaboration with Amgen to understand the aggregation and viscosity of protein solutions in order to develop formulations that are both stable and easily administered.Systems Biology
The "cellular mesoscale" is a range of scales larger than biomolecules but smaller than organelles. This scale contains complex regulatory networks involving hundreds of genes and molecules. Malfunction of these networks leads to devastating diseases like cancer and neurodegeneration. While emergent properties must exist in the mesoscale, we do not know what they are and, therefore, we have a significant gap in understanding how cells function. We take a top-down approach to closing this "mesoscale conceptual gap" using analytic models of experimental data.